Highlight
Shotgun DNA Mapping
Achievement/Results
Larry Herskowitz, advised by Prof. Steven Koch at the Center for High Technology Materials at the University of New Mexico, is working on a new way to map locations within a genome. Through funding from NSF’s Integrated Graduate Education and Research Traineeship program on Integrating Nanotechnology with Cell Biology and Neuroscience, this interdisciplinary project brings together physicists, engineers, and biologists.
The forces that hold the double helical structure of DNA together are sequence-dependent. This is the principal idea behind the shotgun DNA mapping (SDM). The first step in this process is to digest a particular genome using an endonuclease. For this example, the yeast genome will be digested by XhoI. This creates approximately 2,700 different fragments of DNA. Since the yeast genome is published, all of these possible fragments’ sequences are known. This allows for simulation of the unzipping forces for each one of these fragments.
All of these simulations are then combined to form a “library”. With the library complete, a random fragment from the digested genome is chosen and unzipped using an optical tweezer. An optical tweezer is a device that uses a focused laser beam to apply piconewton forces to small objects. In this experiment it is used to unzip DNA and probe the forces that hold the DNA together. These experimental unzipping data are then compared to every entry in the simulated library. One simulation best matches the experimental data. That simulation corresponds to the sequence of the fragment that was unzipped. This allows the researchers to know which fragment was unzipped.
The advantages to this method are that the genome can be taken from living cells. They do not have to be cloned or specially modified for this process to work, which is a large hurdle for most biology labs. Having a mapping technique that circumvents this problem is a large benefit. The highlight is that the simulation for the unzipping forces has been created using a quasi-equilibrium model. Also, this process has been shown to work by using pBR322 experimental data, and hiding them in the yeast simulation library. For over 30 different experimental data sets, SDM was able to find the correct simulation each time.
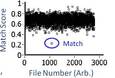
Figures from this proof of principle experiment are attached. Fig. 1 is the comparison of a simulated pBR322 compared to an experimental. Fig. 2 clearly shows the correct simulation standing out from the noise. Finally, Fig. 3 shows a histogram of all 30 different trials of SDM. The overlap of the histograms represents possible false positives, shown to be very small.
Address Goals
The primary goal is to develop a new mapping technique using the binding forces of DNA. Shotgun DNA Mapping does precisely that. This procedure will be able to map loci on genomes without the use of an ensemble averaging method, which loses precious temporal and spatial resolution. In its secondary goal, SDM can be applied to many research areas such as telomere structure, nucleosome remodeling, and alternative splicing studies, thus offering a new experimental tool for a variety of genomic applications.